TR&D3 Research Highlights
Modeling with BioNetGen Gives MMBioS Team Insight into How Immune System Decides to Attack — or Let Be
 Friend or Foe?
Friend or Foe?
A mix of computer modeling and laboratory experiments has helped reveal how the body differentiates “friend from foe.” Using their BioNetGen computer tool for simulating biochemistry, MMBioS members and colleagues have painted a sharper picture of how T cells, the advance scouts of the immune system, decide when to protect bodily tissues from immune attack—and when to lead the attack. The finding may guide future efforts to control human diseases like diabetes and cancer.
When T cells—a type of white blood cell—encounter other cells in the body, “they have to decide what kind of a response to make,” says James Faeder, project co-leader of MMBioS’s Technology Research and Development Project 2 Team and associate professor of computational & systems biology, University of Pittsburgh. “Is that a threat, or is it something benign? Depending on their assessment of a possible threat they can either become activated, immune-boosting cells that kill pathogens, or they can tamp down those responses.”
See more about MMBioS research in cell modeling.
Improved Sampling of Cell-Scale Models using the Weighted Ensemble Strategy
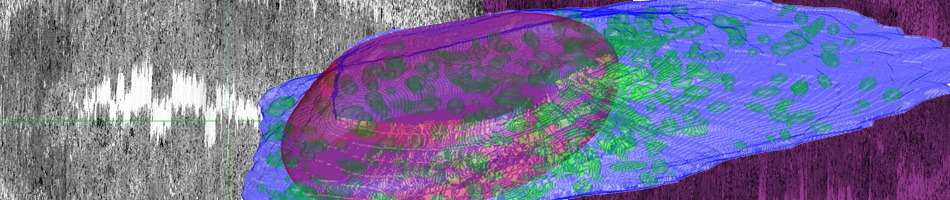
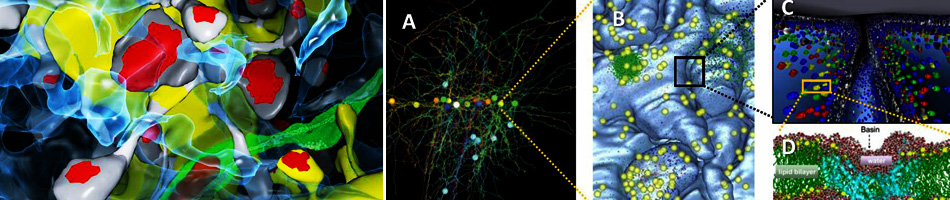
 The “weighted ensemble” (WE) strategy for orchestrating a large set of parallel simulations has been established as an effective tool for efficiently calculating kinetic and equilibrium observables in molecular systems – and now has been extended to spatially resolved cell-scale systems by MMBioS researchers. In a collaboration among several MMBioS groups, the WESTPA implementation of WE has been used to control MCell simulations, including models built using a BioNetGen-CellOrganizer pipeline for situating complex biochemistry within spatially realistic cell models. As sketched, the WE strategy enables computational effort to be focused on rare, difficult-to-sample events – essentially increasing the precision of measurements in the tails of distributions.
The “weighted ensemble” (WE) strategy for orchestrating a large set of parallel simulations has been established as an effective tool for efficiently calculating kinetic and equilibrium observables in molecular systems – and now has been extended to spatially resolved cell-scale systems by MMBioS researchers. In a collaboration among several MMBioS groups, the WESTPA implementation of WE has been used to control MCell simulations, including models built using a BioNetGen-CellOrganizer pipeline for situating complex biochemistry within spatially realistic cell models. As sketched, the WE strategy enables computational effort to be focused on rare, difficult-to-sample events – essentially increasing the precision of measurements in the tails of distributions.  In an application to an MCell model of the frog neuro-muscular junction (NMJ), WE simulation enabled calculation of vesicle release probabilities under particularly challenging low-calcium concentrations that could not be well sampled by conventional MCell simulation: see red-circled points in the figure. This in turn, enabled validation of the model by confirmation of an established NMJ empirical power-law relationship between calcium-concentration and release probability. WE yielded estimates of observables in less overall computing time than would be required in ordinary parallelization, thus exhibiting super-linear parallel performance.
In an application to an MCell model of the frog neuro-muscular junction (NMJ), WE simulation enabled calculation of vesicle release probabilities under particularly challenging low-calcium concentrations that could not be well sampled by conventional MCell simulation: see red-circled points in the figure. This in turn, enabled validation of the model by confirmation of an established NMJ empirical power-law relationship between calcium-concentration and release probability. WE yielded estimates of observables in less overall computing time than would be required in ordinary parallelization, thus exhibiting super-linear parallel performance.
Donovan RM, Tapia JJ, Sullivan DP, Faeder JR, Murphy RF, Dittrich M, Zuckerman DM (2016). Unbiased Rare Event Sampling in Spatial Stochastic Systems Biology Models Using a Weighted Ensemble of Trajectories PLoS Comput Biol. 12(2):e1004611
See more about MMBioS research in molecular modeling
Development and improvements to MCell
The MCell modeling and simulation platform provides the core capabilities for spatial simulation reaction-diffusion dynamics at complex biological interfaces. Highlights of progress include:
libMCell API We have developed a high-level set of routines, which enable the creation and simulation of MCell models via API calls alone, i.e., without invoking any parsed MDL description of the model. Current functionality includes creation of species, defining reactions, defining molecule releases, creating model geometry, and simulation and runtime control.
MCell testing framework This framework, called nutmeg (https://github.com/haskelladdict/nutmeg), has completely replaced our previous, python based unit test framework. During the year we made nutmeg much more robust and added many additional test cases.
Implementation of new simulation capabilities that extend the range of simulations that can be performed in order to address important biological questions, such as the how membrane dynamics and signal transduction interact or how multisite phosphorylation, binding, and cooperativity affect signaling. Highlights include:
- Dynamic geometries Users can now create dynamical meshes using Blender and use these to drive an MCell simulation of reaction and diffusion with moving boundaries.
- Modeling pipeline Users can build complex biochemical models using BioNetGen and sample complex cellular geometries from imaging data using CellOrganizer and combine those into a single model that can be simulated in CellBlender (in collaboration with TRD3).
- Recovery of protein-protein interactions and other aspect of biochemical mechanisms from reaction network models using Atomizer. The automated conversion of these models into a rule-based format allows for comparative analysis of models and reuse of existing model components.
- Visualization of regulatory structurein rule-based models New visualization methods have been developed that enable global visualization of models exhibiting combinatorial complexity.
D. P. Sullivan, J. J. Tapia, R. Arepally, J. Czech, R. F. Murphy, M. Dittrich, and J. R. Faeder, “Design Automation for Biological Models : A Pipeline that Incorporates Spatial and Molecular Complexity,” in 25th edition of the Great Lakes Symposium on VLSI, 2015, pp. 321–323.
J. A. P. Sekar, J.-J. Tapia, and J. R. Faeder, “Visualizing Regulation in Rule-based Models.” Submitted to Bioinformatics . arXiv:1509.00896 [q-bio.QM].
See more about MMBioS research in cell modeling.
Development of CellBlender
CellBlender is a graphical interface for model construction, simulation, and analysis of complex spatial models of reaction-diffusion systems Highlights of progress include
- A major redesign of the interface to consolidate functionality based on a single toolbar.
- Implementation of a robust parameter handling system
- Functionality for running simulations from within the interface taking advantage of parallel computing resources allowing users to manage on-going simulation streams.
- Direct access to run-time error logs within the Blender interface.
- Model and geometry importers that allow import of compartmental reaction network models in either SBML or compartmental BNGL format.
- Implementation of internal data model provides backward compatibility with future CellBlender versions.
- Release of CellBlender version 1.0 and publication of an article featuring MCell and CellBlender in the Encyclopedia of Computational Neuroscience
T. Bartol, M. Dittrich, and J. Faeder, “MCell,” in Encyclopedia of Computational Neuroscience, D. Jaeger and R. Jung, Eds. Springer New York, 2014, pp. 1–5.
See more about MMBioS research in cell modeling.
Stochastic Simulation
 Systems biology, the quantitative study of complex interacting biological systems, is becoming increasingly demanding of computational resources. As models become able to capture truly physiological behavior, some of the most difficult computations may also be the most important, such as a rare transition from normal to pathological behavior.
Systems biology, the quantitative study of complex interacting biological systems, is becoming increasingly demanding of computational resources. As models become able to capture truly physiological behavior, some of the most difficult computations may also be the most important, such as a rare transition from normal to pathological behavior.
A novel approach to meet growing computational demands is described in a paper recently accepted to the Journal of Chemical Physics ("Efficient Stochastic Simulation of Chemical Kinetics Networks Using A Weighted Ensemble Of Trajectories"). First-author Donovan, in collaboration with MMBioS investigators Faeder and Zuckerman, applied a sophisticated parallel simulation algorithm originally developed for small-scale molecular systems to systems biology models.
The results were eye-catching: the new approach was more computationally efficient at modeling rare events than standard methods by orders of magnitude, even for a very complex model with thousands of reacting species.
Donovan RM, Sedgewick AJ, Faeder JR, Zuckerman DM (2013) Efficient Stochastic Simulation of Chemical Kinetics Networks Using A Weighted Ensemble Of Trajectories J. Chem. Phys. 139:115105.
TR&D2 Research Highlights
Cell Modeling Research Highlights

Pre-post synaptic alignment through neuroligin-1 tunes synaptic transmission efficiency
TR&D2 investigators and collaborators describe organizing role of neuroligin-1 to align post-synaptic AMPA Receptors with pre-synaptic release sites into trans-synaptic “nano-columns” to enhance signaling.(Read more)

BioNetGen Gives Insight into Immune System
A mix of modeling with BioNetGen and laboratory experiments has painted a sharper picture of how T cells decide when to protect the body from immune attack —and when to lead the attack. Read more
 Memory Capacity of Brain 10 Times More than Previously Thought
Memory Capacity of Brain 10 Times More than Previously Thought
MMBioS members Terry Sejnowski and Tom Bartol, along with other Salk Institute collaborators, have achieved critical insight into the size of neural connections, putting the memory capacity of the brain far higher than common estimates. Read more
 Improved Sampling of Cell-Scale Models using the Weighted Ensemble Strategy
Improved Sampling of Cell-Scale Models using the Weighted Ensemble Strategy
The “weighted ensemble” (WE) strategy for orchestrating a large set of parallel simulations has been established as an effective tool for efficiently calculating kinetic and equilibrium observables in molecular systems – and now has been extended to spatially resolved cell-scale systems by MMBioS researchers. Read more
Development and improvements to MCell
MCell is a modeling and simulation platform which provides the core capabilities for spatial simulation reaction-diffusion dynamics at complex biological interfaces. Recent improvements include libMCell, an MCell testing framework and new simulation capabilities that extend the range of simulations that can be performed. Read more
CellBlender is a graphical interface for model construction, simulation, and analysis of complex spatial models of reaction-diffusion systems. The interface has been redesigned and functionality expanded. Read more
 Synaptic Facilitation Revealed
Synaptic Facilitation Revealed
An investigation of several mechanisms of short-term facilitation at the frog neuromuscular junction concludes that the presence of a second class of calcium sensor proteins distinct from synaptotagmin can explain known properties of facilitation. Read more

Synapses are the connections between neurons in the brain or between neurons and muscle fibers...read more.
TR&D4 Research Highlights
Image Processing & Analysis Research Highlights

Tools for determining the spatial relationships
An important task for understanding how cells are organized is determining which components have spatial patterns that are related to each other. Read more

Pipeline for creation of spatiotemporal maps
Using a combination of diffeomorphic methods and improved cell segmentation, we developed a CellOrganizer pipeline for use in DPB4 to construct models of the 4D distributions of actin and 8 of its regulators during the response of T cells to antigen presentation. Read more

Development of models of cell and nuclear shape
We added a major new capability to our open source CellOrganizer system, the ability to construct diffeomorphic models of cell and nuclear shape. The models were developed because of the needs of DBP4... Read More

Anatomy and Function of an Excitatory Network in the Visual Cortex
MMBioS researcher Greg Hood’s collaboration with Wei-Chung Allen Lee of Harvard University and R. Clay Reid of the Allen Institute for Brain Science concerning the reconstruction of an excitatory nerve-cell network in the mouse brain cortex at a subcellular level using the AlignTK software has been published in Nature. Read more
 Cellorganizer 2.0 Major Release
Cellorganizer 2.0 Major Release
A major new release of the CellOrganizer system for creating image-derived models of cell shape and organization has just been published. Read more.
TR&D1 Research Highlights
Molecular Modeling Research Highlights

Monoamine transporters: structure, intrinsic dynamics and allosteric regulation/strong><
T&RD1 investigators Mary Cheng and Ivet Bahar published an invited review article in Nature Structural & Molecular Biology, addressing recent progress in the elucidation of the structural dynamics of MATs and their conformational landscape and transitions, as well as allosteric regulation mechanisms. (Read more)

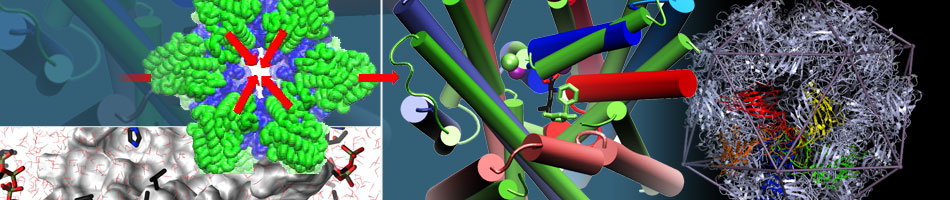
Trimerization of dopamine transporter triggered by AIM-100 binding
The Bahar (TR&D1) and Sorkin (DBP3) labs explored the trimerization of dopamine transporter (DAT) triggered by a furopyrimidine, AIM-100, using a combination of computational and biochemical methods, and single-molecule live-cell imaging assays. (Read more)

Our findings highlight an important mechanism by which proteins genetically implicated in Parkinson’s disease (PD; PINK1) and frontotemporal dementia (FTD; VCP) interact to support the health and maintenance of neuronal arbors.(Read more)

New tool to predict pathogenicity of missense variants based on structural dynamics: RHAPSODY
We demonstrated that the analysis of a protein’s intrinsic dynamics can be successfully used to improve the prediction of the effect of point mutations on a protein functionality. This method employs ANM/GNM tools (Read more)

Multi-scale Hybrid Methodology
The hybrid methodology, coMD, that we have recently developed [1] has been recently extended to construct the energy landscape near the functional states of LeuT (Fig 1) [2]. This is the first energy landscape constructed for this NSS family member. Read more

Insights into the cooperative dynamics of AMPAR
Comparative analysis of AMPAR and NMDAR dynamics reveals striking similiarities, opening the way to designing new modulators of allosteric interactions. Read more

New Release of the iGNM Database
We have updated our iGNM database. The updated iGNM 2.0 covers more than 95% of the structures in the Protein Data Bank. Read more
 Sparse Graphical Models of Protein:Protein Interactions
Sparse Graphical Models of Protein:Protein Interactions
DgSpi is a new method for learning and using graphical models that explicitly represent the amino acid basis for interaction specificity and extend earlier classification-oriented approaches to predict ΔG of binding. Read more
 Improved Sampling of Cell-Scale Models using the WE Strategy
Improved Sampling of Cell-Scale Models using the WE Strategy
The WE strategy for orchestrating a large set of parallel simulations has now been extended to spatially resolved cell-scale systems. The WESTPA implementation of WE has been used to control MCell simulations, including models built using a BioNetGen-CellOrganizer pipeline for situating complex biochemistry within spatially realistic cell models. Read more
 Molecular Mechanism of Dopamine Transport by hDAT
Molecular Mechanism of Dopamine Transport by hDAT
Dopamine transporters (DATs) control neurotransmitter dopamine (DA) homeostasis by reuptake of excess DA, assisted by sodium and chloride ions. The recent resolution of DAT structure (dDAT) from Drosophila permits us for the first time to directly view the sequence of events involved in DA reuptake in human DAT (hDAT). Read more
 Advancing Parallel Bio-simulations
Advancing Parallel Bio-simulations
A new non-Markovian analysis can eliminate bias in estimates of long-timescale behavior, such as the mean first-passage time for the dissociation of methane molecules in explicit solvent. Read more
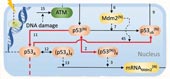
Controlling ionizing radiation (IR)-induced cell death mitigates radiation damage. Examining tumor suppressor protein p53 network dynamics in response to IR damage found that the strength of p53 transcriptional activity and its coupling (or timing with respect) to mitochondrial pore opening are major determinants of cell fate. Read more
Our recent study highlights the role of the helical hairpin HP2 as an intracellular gate, in addition to its role as an extracellular gate. Read more.
Unraveling the molecular mechanism of function of NSS family members has been a challenge due to the involvement of both local (EC or IC gate opening/closure) and global (between outward- and inward-facing) changes in structure. Read more
Latest News
Upcoming Events
| No events |
Past Events
| No events |
Collaborate with MMBioS
Research Highlights
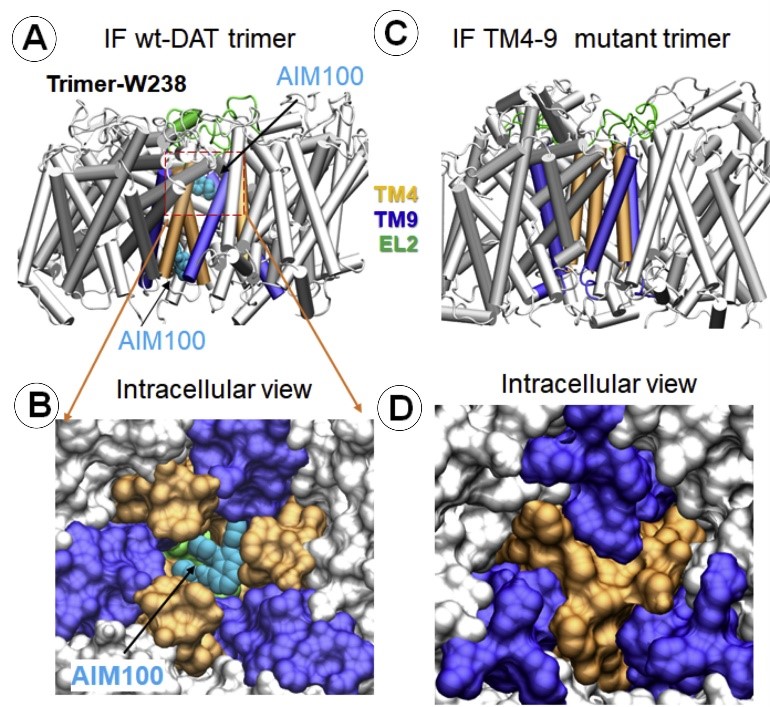
The Bahar (TR&D1) and Sorkin (DBP3) labs published an article in the Journal of Biological Chemistry, selected as one of JBC's "Editors' Picks. Our results demonstrate a direct coupling between conformational dynamics of DAT, functional activity of the transporter and its oligomerization leading to endocytosis. The high specificity of such coupling for DAT makes the TM4-9 hub a new target for pharmacological modulation of DAT activity and subcellular localization. (Read more)

Differences in the intrinsic spatial dynamics of the chromatin contribute to cell differentiation
Comparison with RNA-seq expression data reveals a strong overlap between highly expressed genes and those distinguished by high mobilities in the present study, in support of the role of the intrinsic spatial dynamics of chromatin as a determinant of cell differentiation. (Read more)

Nanoscale co-organization and coactivation of AMPAR, NMDAR, and mGluR at excitatory synapses
Work by TR&D2 Investigators and collaborators provide insights into the nanometer scale organization of postsynaptic glutamate receptors using a combination of dual-color superresolution imaging, electrophysiology, and computational modeling. (Read more)

Parallel Tempering with Lasso for model reduction in systems biology
TR&D3 Investigators and collaborators develop PTLasso, a Bayesian model reduction approach that combines Parallel Tempering with Lasso regularization, to automatically extract minimal subsets of detailed models that are sufficient to explain experimental data. On both synthetic and real biological data, PTLasso is an effective method to isolate distinct parts of a larger signaling model that are sufficient for specific data. (Read more)

Image-derived models of cell organization changes during differentiation and drug treatments
Our work on modeling PC12 cells undergoing differentiation into neuron-like morphologies (under C&SP11, completed) has been published in Molecular Biology of the Cell. We have also made the large dataset of 3D images collected in that study available through Dryad. (Read more)

Monoamine transporters: structure, intrinsic dynamics and allosteric regulation
T&RD1 investigators Mary Cheng and Ivet Bahar published an invited review article in Nature Structural & Molecular Biology, addressing recent progress in the elucidation of the structural dynamics of MATs and their conformational landscape and transitions, as well as allosteric regulation mechanisms. (Read more)

Trimerization of dopamine transporter triggered by AIM-100 binding
The Bahar (TR&D1) and Sorkin (DBP3) labs explored the trimerization of dopamine transporter (DAT) triggered by a furopyrimidine, AIM-100, using a combination of computational and biochemical methods, and single-molecule live-cell imaging assays. (Read more)

Pre-post synaptic alignment through neuroligin-1 tunes synaptic transmission efficiency
TR&D2 investigators and collaborators describe organizing role of neuroligin-1 to align post-synaptic AMPA Receptors with pre-synaptic release sites into trans-synaptic “nano-columns” to enhance signaling.(Read more)

Inferring the Assembly Network of Influenza Virus
In an article in PLoS Computational Biology, MMBioS TR&D4 members Xiongto Ruan and Bob Murphy collaborated with Seema Lakdawala to address this question of the assembly network of the Influenza virus.(Read more)

Our findings highlight an important mechanism by which proteins genetically implicated in Parkinson’s disease (PD; PINK1) and frontotemporal dementia (FTD; VCP) interact to support the health and maintenance of neuronal arbors.(Read more)

Improved methods for modeling cell shape
In a recent paper in Bioinformatics, Xiongtao Ruan and Bob Murphy of TR&D4 addressed the question of how best to model cell and nuclear shape.(Read more)

New tool to predict pathogenicity of missense variants based on structural dynamics: RHAPSODY
We demonstrated that the analysis of a protein’s intrinsic dynamics can be successfully used to improve the prediction of the effect of point mutations on a protein functionality. This method employs ANM/GNM tools (Read more)

New method for investigating chromatin structural dynamics.
By adapting the Gaussian Network Model (GNM) protein-modeling framework, we were able to model chromatin dynamics using Hi-C data, which led to the identification of novel cross-correlated distal domains (CCDDs) that were found to also be associated with increased gene co-expression. (Read more)

Structural elements coupling anion conductance and substrate transport identified
We identified an intermediate anion channeling state (iChS) during the global transition from the outward facing (OF) to inward facing state (IFS). Our prediction was tested and validated by experimental study conducted in the Amara lab (NIMH). Critical residues and interactions were analyzed by SCAM, electrophysiology and substrate uptake experiments (Read more)

Integrating MMBioS technologies for multiscale discovery
TR&D teams driven by individual DBPs are naturally joining forces, integrating their tools to respond to the needs of the DBP, and creating integrative frameworks for combining structural and kinetic data and computing technologies at multiple scales. (Read more)

Large scale visualization of rule-based models.
Signaling in living cells is mediated through a complex network of chemical interactions. Current predictive models of signal pathways have hundreds of reaction rules that specify chemical interactions, and a comprehensive model of a stem cell or cancer cell would be expected to have many more. Visualizations of rules and their interactions are needed to navigate, organize, communicate and analyze large signaling models. (Read more)

Integration of MCellR into MCell/CellBlender
Using spatial biochemical models of SynGAP/PSD95, MMBioS investigators were able to merge the MCellR code-base with the MCell code-base and validate its utility and correctness of this sophisticated technology now easily accessible through the MCell/CellBlender GUI. (Read more)
Causal relationships of spatial distributions of T cell signaling proteins
The idea is to identify a relationship in which a change in the concentration of one protein in one cell region consistently is associated with a change in the concentration of another protein in the same or a different region. We used the data from our Science Signaling paper reported last year to construct a model for T cells undergoing stimulation by both the T cell receptor and the costimulatory receptor. (Read more...)

BioNetGen modeling helps reveal immune system response decision
To attack or to let be is an important decision that our immune systems must make to protect our bodies from foreign invaders or protect bodily tissues from an immune attack. Using modeling and experiments, we have painted a sharper picture of how T cells make these critical decisions. (Read more)

Tools for determining the spatial relationships between different cell components
An important task for understanding how cells are organized is determining which components have spatial patterns that are related to each other.Read more

Pipeline for creation of spatiotemporal maps
Using a combination of diffeomorphic methods and improved cell segmentation, we developed a CellOrganizer pipeline for use in DPB4 to construct models of the 4D distributions of actin and 8 of its regulators during the response of T cells to antigen presentation. Read more

Multi-scale Hybrid Methodology
The hybrid methodology, coMD, that we have recently developed [1] has been recently extended to construct the energy landscape near the functional states of LeuT (Fig 1) [2]. This is the first energy landscape constructed for this NSS family member. Read more

Insights into the cooperative dynamics of AMPAR
Comparative analysis of AMPAR and NMDAR dynamics reveals striking similarities, opening the way to designing new modulators of allosteric interactions. Read more

Improved Sampling of Cell-Scale Models using the WE Strategy
The WE strategy for orchestrating a large set of parallel simulations has now been extended to spatially resolved cell-scale systems. The WESTPA implementation of WE has been used to control MCell simulations, including models built using a BioNetGen-CellOrganizer pipeline for situating complex biochemistry within spatially realistic cell models. Read more

Anatomy and Function of an Excitatory Network in the Visual Cortex
MMBioS researcher Greg Hood’s collaboration with Wei-Chung Allen Lee of Harvard University and R. Clay Reid of the Allen Institute for Brain Science concerning the reconstruction of an excitatory nerve-cell network in the mouse brain cortex at a subcellular level using the AlignTK software has been published in Nature. Read more
 Molecular Mechanism of Dopamine Transport by hDAT
Molecular Mechanism of Dopamine Transport by hDAT
Dopamine transporters (DATs) control neurotransmitter dopamine (DA) homeostasis by reuptake of excess DA, assisted by sodium and chloride ions. The recent resolution of DAT structure (dDAT) from Drosophila permits us for the first time to directly view the sequence of events involved in DA reuptake in human DAT (hDAT). Read more
 Synaptic Facilitation Revealed
Synaptic Facilitation Revealed
An investigation of several mechanisms of short-term facilitation at the frog neuromuscular junction concludes that the presence of a second class of calcium sensor proteins distinct from synaptotagmin can explain known properties of facilitation. Read more
 Sparse Graphical Models of Protein:Protein Interactions
Sparse Graphical Models of Protein:Protein Interactions
DgSpi is a new method for learning and using graphical models that explicitly represent the amino acid basis for interaction specificity and extend earlier classification-oriented approaches to predict ΔG of binding. Read more
 Advancing Parallel Bio-simulations
Advancing Parallel Bio-simulations
A new non-Markovian analysis can eliminate bias in estimates of long-timescale behavior, such as the mean first-passage time for the dissociation of methane molecules in explicit solvent. Read more